# Define earth-tone palette inspired by palm habitats
# (forest floor browns, trunk grays, canopy greens)
palette_palms <- c(
"Nypoideae" = "#8B7355", # Mangrove mud brown
"Arecoideae" = "#4A6741", # Understory green
"Coryphoideae" = "#B8956A", # Sandy tan (arid habitats)
"Ceroxyloideae" = "#6B8E8F", # Mountain mist blue-gray
"Calamoideae" = "#5C4033" # Dark rattan brown
)
# Get tallest species per subfamily for annotation
tallest_species <- size_data %>%
group_by(palm_subfamily) %>%
slice_max(max_stem_height_m, n = 1) %>%
ungroup() %>%
mutate(
label = paste0(acc_genus, " ", acc_species, "\n", round(max_stem_height_m, 0), "m")
)
# Create the plot
ggplot(size_data, aes(x = palm_subfamily, y = max_stem_height_m, fill = palm_subfamily)) +
# Violin plot for distribution shape
geom_violin(
alpha = 0.6,
trim = FALSE,
scale = "width",
adjust = 1.2
) +
# Boxplot overlay for summary statistics
geom_boxplot(
width = 0.15,
alpha = 0.8,
outlier.alpha = 0.4,
outlier.size = 1,
color = "gray20"
) +
# Add reference lines for stratification zones
geom_hline(yintercept = 5, linetype = "dashed", color = "gray40", linewidth = 0.4) +
geom_hline(yintercept = 15, linetype = "dashed", color = "gray40", linewidth = 0.4) +
# Annotate stratification zones
annotate("text", x = 0.6, y = 2.5, label = "Understory",
hjust = 0, size = 3, color = "gray30", fontface = "italic") +
annotate("text", x = 0.6, y = 10, label = "Midstory",
hjust = 0, size = 3, color = "gray30", fontface = "italic") +
annotate("text", x = 0.6, y = 25, label = "Canopy",
hjust = 0, size = 3, color = "gray30", fontface = "italic") +
# Label tallest species (excluding Nypoideae with 0m)
geom_text_repel(
data = tallest_species %>% filter(max_stem_height_m > 0),
aes(label = label),
size = 2.8,
fontface = "italic",
color = "gray20",
nudge_x = 0.3,
segment.color = "gray50",
segment.size = 0.3,
min.segment.length = 0
) +
# Color and styling
scale_fill_manual(values = palette_palms) +
scale_y_continuous(
breaks = seq(0, 180, 20),
limits = c(0, 180),
expand = c(0, 0)
) +
coord_flip() +
labs(
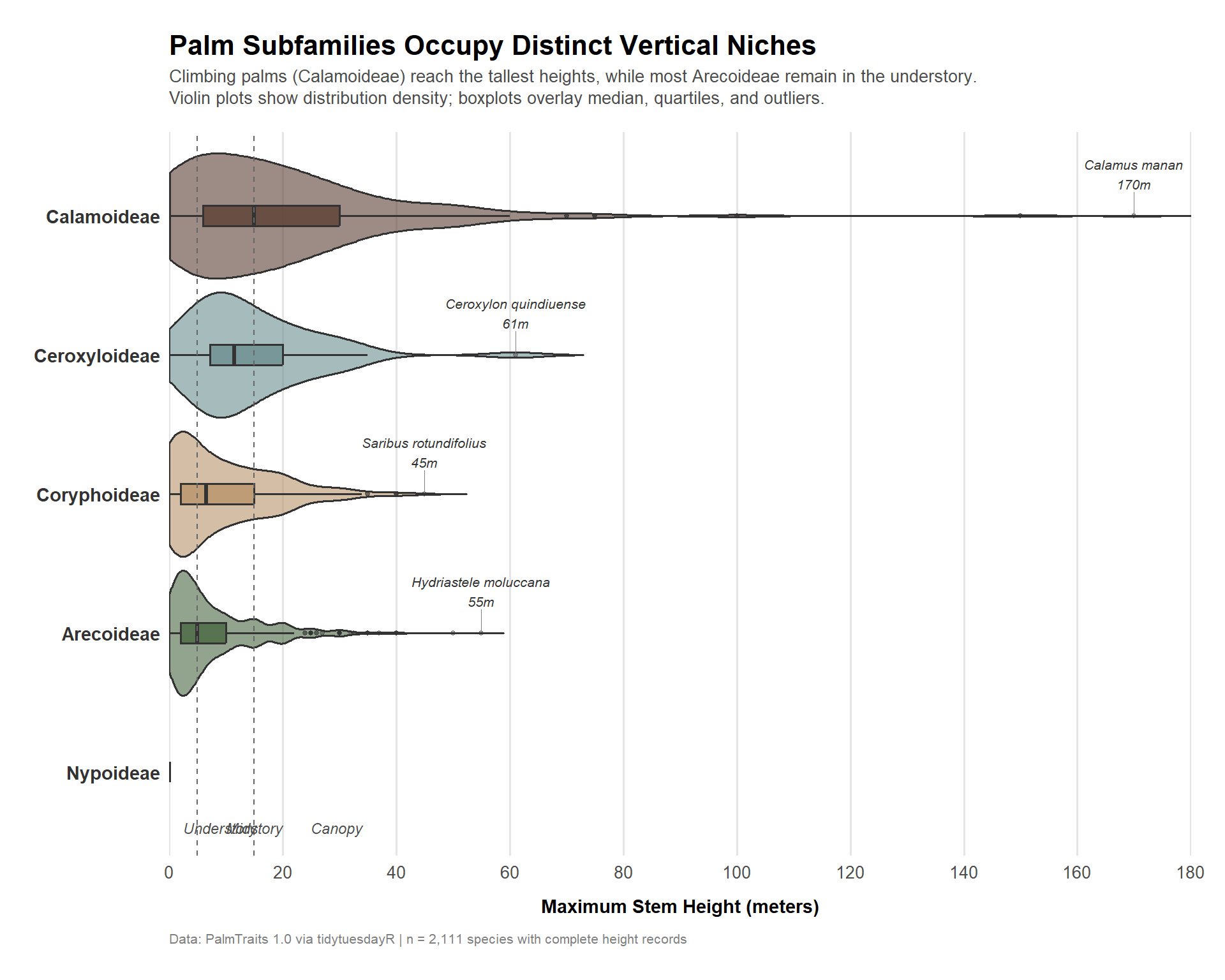
title = "**Palm Subfamilies Occupy Distinct Vertical Niches**",
subtitle = "Climbing palms (Calamoideae) reach the tallest heights, while most Arecoideae remain in the understory.<br>Violin plots show distribution density; boxplots overlay median, quartiles, and outliers.",
x = NULL,
y = "Maximum Stem Height (meters)",
caption = "Data: PalmTraits 1.0 via tidytuesdayR | n = 2,111 species with complete height records"
) +
theme_minimal(base_size = 12) +
theme(
legend.position = "none",
plot.title = element_markdown(size = 16, face = "bold", margin = margin(b = 5)),
plot.subtitle = element_markdown(size = 10, color = "gray30", lineheight = 1.3, margin = margin(b = 15)),
plot.caption = element_text(size = 8, color = "gray50", hjust = 0, margin = margin(t = 10)),
axis.text.y = element_text(size = 11, face = "bold", color = "gray20"),
axis.text.x = element_text(size = 10),
axis.title.x = element_text(size = 11, face = "bold", margin = margin(t = 10)),
panel.grid.major.y = element_blank(),
panel.grid.minor = element_blank(),
panel.grid.major.x = element_line(color = "gray90"),
plot.margin = margin(20, 20, 20, 20)
)