# Convert to sf for spatial plotting
site_sf <- site_stats %>%
st_as_sf(coords = c("longitude", "latitude"), crs = 4326)
# Get Australia coastline for map context
aus <- ne_countries(scale = "large", country = "Australia", returnclass = "sf")
# Grade colors from Hiroshige — clean blues to contaminated reds
grade_colors <- setNames(
as.character(hiroshige[c(8, 6, 3, 1)]),
c("Good", "Fair", "Poor", "Very Poor")
)
# Split sites: coastal/harbour vs western Sydney rivers
coastal_sf <- site_sf %>% filter(region != "Western Sydney")
western_sf <- site_sf %>% filter(region == "Western Sydney")
# -- Coastal panel -- harbour & ocean beaches
coastal_bbox <- st_bbox(coastal_sf)
c_pad <- 0.02
c_xlim <- c(
as.numeric(coastal_bbox["xmin"]) - c_pad,
as.numeric(coastal_bbox["xmax"]) + c_pad
)
c_ylim <- c(
as.numeric(coastal_bbox["ymin"]) - c_pad,
as.numeric(coastal_bbox["ymax"]) + c_pad
)
# Fetch harbour waterways from OSM (tryCatch for API resilience)
coast_osm_bbox <- c(c_xlim[1], c_ylim[1], c_xlim[2], c_ylim[2])
coast_river_lines <- tryCatch(
{
q <- opq(bbox = coast_osm_bbox, timeout = 60) %>%
add_osm_feature(key = "waterway", value = c("river", "canal")) %>%
osmdata_sf()
q$osm_lines
},
error = function(e) NULL
)
p_coast <- ggplot() +
geom_sf(data = aus, fill = "grey90", color = "grey50", linewidth = 0.3) +
{
if (!is.null(coast_river_lines) && nrow(coast_river_lines) > 0) {
geom_sf(
data = coast_river_lines,
color = "#4a90b8",
linewidth = 0.4,
alpha = 0.7
)
}
} +
geom_sf(
data = coastal_sf,
aes(color = quality_grade, size = p90_entero),
alpha = 0.75
) +
coord_sf(xlim = c_xlim, ylim = c_ylim, expand = FALSE) +
scale_color_manual(values = grade_colors, drop = FALSE) +
scale_size_continuous(
range = c(2, 10),
trans = "log10",
breaks = c(20, 100, 500),
limits = range(site_stats$p90_entero, na.rm = TRUE),
labels = scales::comma_format(),
name = "90th Percentile\nEnterococci"
) +
labs(title = "Coastal & Harbour Sites", color = "Quality Grade") +
theme_minimal(base_size = 11) +
theme(
plot.title = element_text(face = "bold", size = 12),
panel.background = element_rect(fill = "#d6eaf8", color = NA),
panel.grid = element_line(color = "grey85"),
axis.text = element_text(size = 7)
)
# -- Western Sydney panel -- fetch rivers from OpenStreetMap
western_bbox_vals <- st_bbox(western_sf)
w_pad <- 0.05
w_xlim <- c(
as.numeric(western_bbox_vals["xmin"]) - w_pad,
as.numeric(western_bbox_vals["xmax"]) + w_pad
)
w_ylim <- c(
as.numeric(western_bbox_vals["ymin"]) - w_pad,
as.numeric(western_bbox_vals["ymax"]) + w_pad
)
# Query OSM for rivers in the Western Sydney area (named rivers only)
osm_bbox <- c(w_xlim[1], w_ylim[1], w_xlim[2], w_ylim[2])
river_lines <- tryCatch(
{
q <- opq(bbox = osm_bbox, timeout = 60) %>%
add_osm_feature(key = "waterway", value = c("river")) %>%
osmdata_sf()
q$osm_lines %>% filter(!is.na(name))
},
error = function(e) NULL
)
p_west <- ggplot() +
geom_sf(data = aus, fill = "grey90", color = "grey50", linewidth = 0.3) +
{
if (!is.null(river_lines) && nrow(river_lines) > 0) {
geom_sf(
data = river_lines,
color = "#4a90b8",
linewidth = 0.5,
alpha = 0.8
)
}
} +
geom_sf(
data = western_sf,
aes(color = quality_grade, size = p90_entero),
alpha = 0.75
) +
coord_sf(xlim = w_xlim, ylim = w_ylim, expand = FALSE) +
scale_color_manual(values = grade_colors, drop = FALSE) +
scale_size_continuous(
range = c(2, 10),
trans = "log10",
breaks = c(20, 100, 500),
limits = range(site_stats$p90_entero, na.rm = TRUE),
labels = scales::comma_format(),
name = "90th Percentile\nEnterococci"
) +
labs(title = "Western Sydney River Sites", color = "Quality Grade") +
theme_minimal(base_size = 11) +
theme(
plot.title = element_text(face = "bold", size = 12),
panel.grid = element_line(color = "grey92"),
axis.text = element_text(size = 7)
)
# Combine maps — standalone version with legend
p_map <- p_coast +
p_west +
plot_layout(widths = c(1.4, 1), guides = "collect") +
plot_annotation(
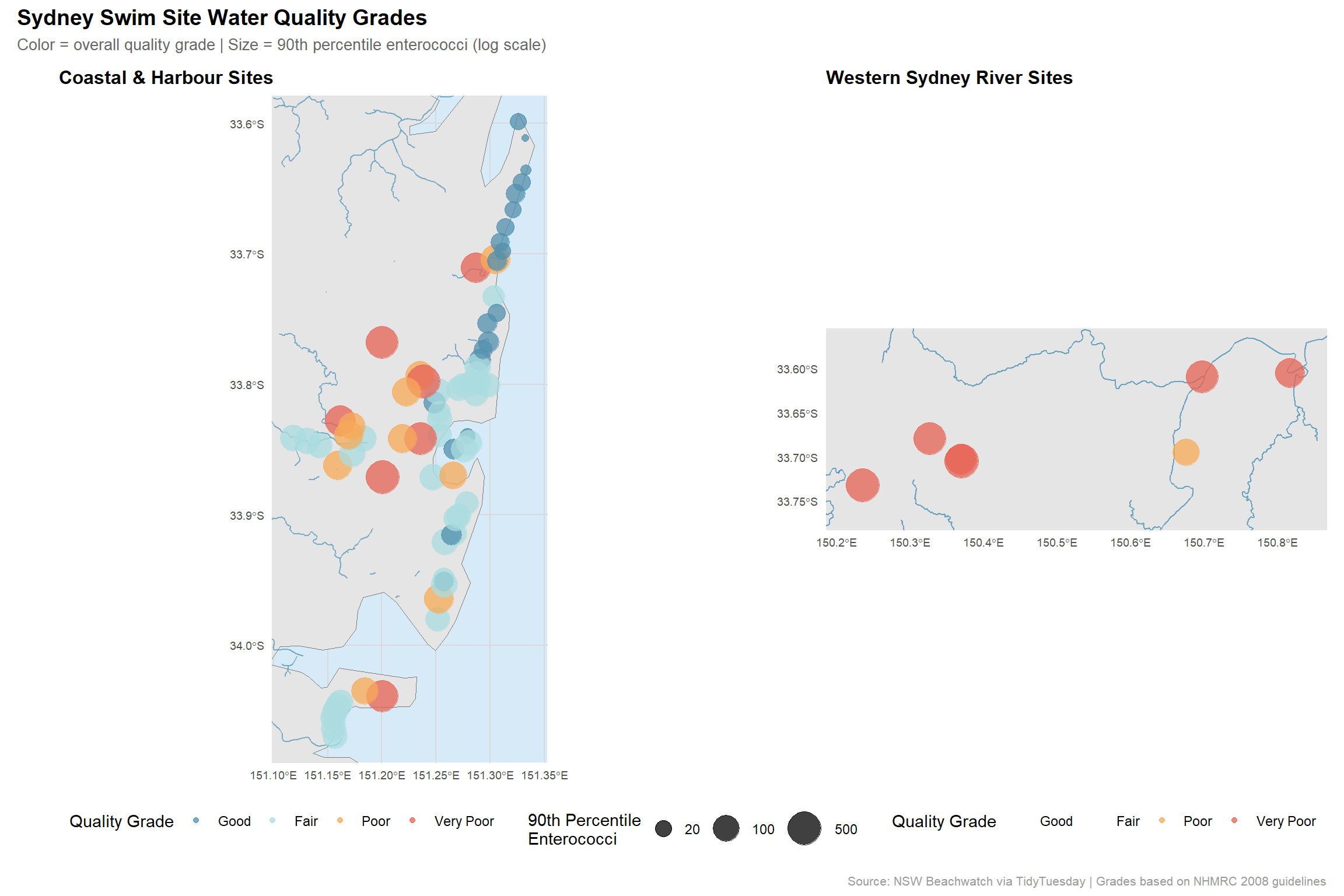
title = "Sydney Swim Site Water Quality Grades",
subtitle = "Color = overall quality grade | Size = 90th percentile enterococci (log scale)",
caption = "Source: NSW Beachwatch via TidyTuesday | Grades based on NHMRC 2008 guidelines",
theme = theme(
plot.title = element_text(face = "bold", size = 14),
plot.subtitle = element_text(color = "grey40", size = 10),
plot.caption = element_text(color = "grey60", size = 8)
)
) &
theme(legend.position = "bottom")
p_map