# Palette: ghibli::MononokeMedium (7 colors — plenty for 6 WHO regions)
# Warm earthy tones from Princess Mononoke: fits the gravity of the topic
region_colors <- paletteer::paletteer_d("ghibli::MononokeMedium", n = 6)
p <- plot_data %>%
ggplot(aes(x = year, y = total_cases, fill = region_label)) +
geom_area(
alpha = 0.92,
color = "white",
linewidth = 0.25,
position = "stack"
) +
# Vertical reference lines for key events
geom_vline(xintercept = 2000, color = "#555555", linetype = "dashed", linewidth = 0.5) +
geom_vline(xintercept = year_peak, color = "#8B1A1A", linetype = "dotted", linewidth = 0.6) +
# Annotations
annotate(
"text",
x = 2001, y = max(global_trend$total_cases, na.rm = TRUE) * 0.92,
label = "Measles Initiative\nlaunched (2001)",
hjust = 0, size = 3.2, color = "#444444", fontface = "italic"
) +
annotate(
"text",
x = year_peak + 0.3,
y = max(global_trend$total_cases, na.rm = TRUE) * 0.92,
label = sprintf("2019 resurgence\n%s reported cases", scales::comma(peak_val)),
hjust = 0, size = 3.2, color = "#8B1A1A", fontface = "bold.italic"
) +
# Scales
scale_x_continuous(breaks = seq(1980, 2025, by = 5), expand = expansion(mult = c(0.01, 0.05))) +
scale_y_continuous(
labels = scales::label_comma(scale = 1e-3, suffix = "K"),
expand = expansion(mult = c(0, 0.05))
) +
scale_fill_manual(values = as.character(region_colors)) +
# Labels
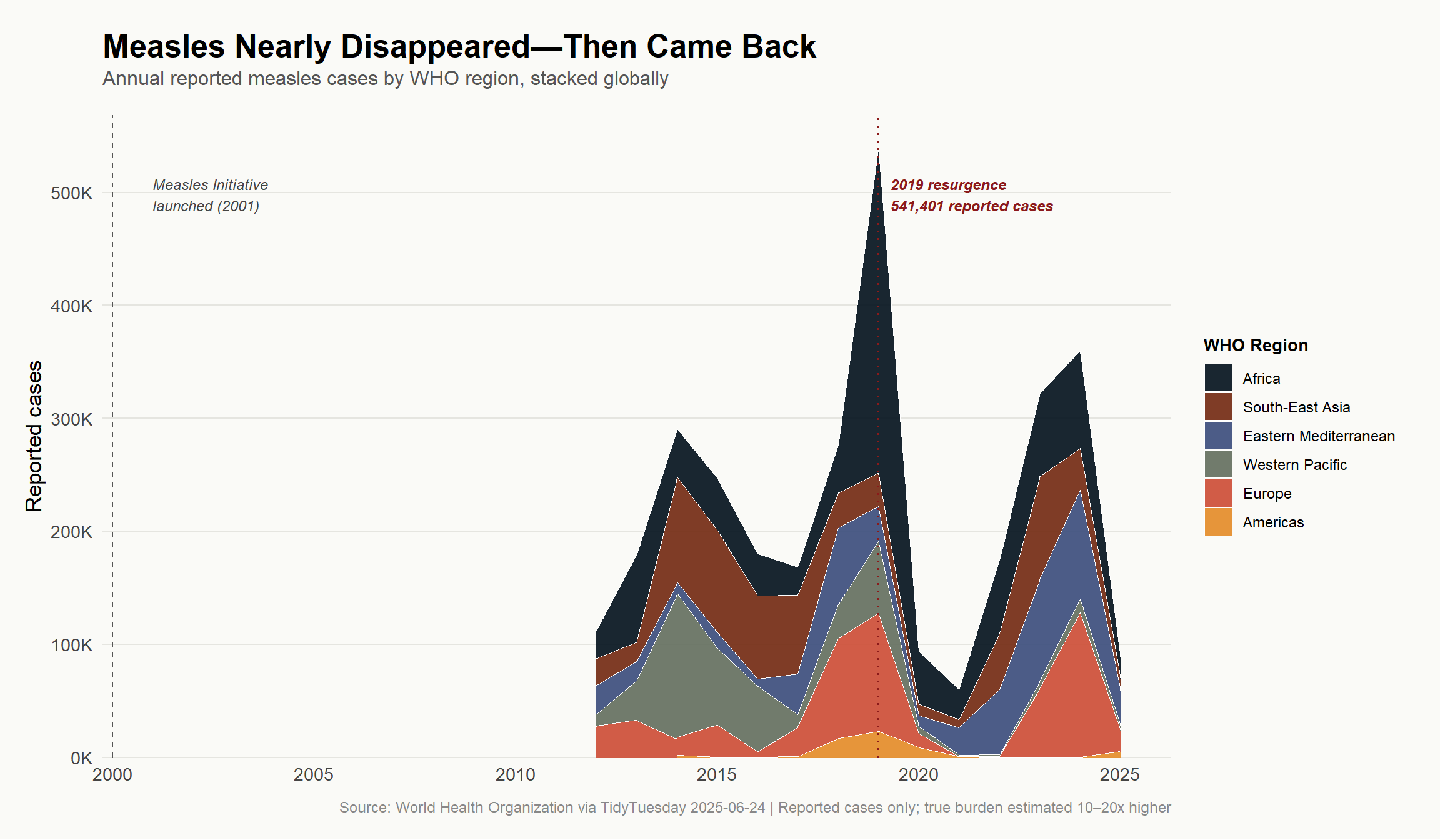
labs(
title = "**Measles Nearly Disappeared—Then Came Back**",
subtitle = "Annual reported measles cases by WHO region, stacked globally",
caption = "Source: World Health Organization via TidyTuesday 2025-06-24 | Reported cases only; true burden estimated 10–20x higher",
x = NULL,
y = "Reported cases",
fill = "WHO Region"
) +
theme_minimal(base_size = 13) +
theme(
plot.title = element_markdown(size = 19, face = "bold", margin = margin(b = 4)),
plot.subtitle = element_text(size = 12, color = "#555555", margin = margin(b = 16)),
plot.caption = element_text(size = 9, color = "#888888", margin = margin(t = 10)),
plot.background = element_rect(fill = "#FAFAF7", color = NA),
panel.background = element_rect(fill = "#FAFAF7", color = NA),
panel.grid.major.x = element_blank(),
panel.grid.minor = element_blank(),
panel.grid.major.y = element_line(color = "#E5E5E0", linewidth = 0.4),
axis.text = element_text(color = "#444444"),
legend.position = "right",
legend.title = element_text(size = 10, face = "bold"),
legend.text = element_text(size = 9),
plot.margin = margin(20, 20, 15, 15)
)
p